In last week’s post we described
a free online tool for predicting bad behavior of compounds in various assays.
But as we noted, you often get what you pay for, and computational methods can’t
(yet) take the place of experimentation. In a new (open-access) J. Med.
Chem. paper, Steven LaPlante and collaborators at NMX and INRS describe a
roadmap for discovering, validating, and advancing weak fragments. They call it
NMR by SAR
Unlike SAR by NMR, the grand-daddy of fragment-finding techniques which involves
protein-detected NMR, NMR for SAR focuses heavily on the ligand. The
researchers illustrate the process by finding ligands for the protein HRAS, for
which drug discovery has lagged in comparison to its sibling KRAS.
The researchers started by
screening the G12V mutant form of HRAS in its inactive (GDP-bound) state. They screened
their internal library of 461 fluorinated fragments in pools of 11-15 compounds
(each at ~0.24 mM) using 19F NMR. An initial screen at 15 µM protein
produced a very low hit rate, so the protein concentration was increased to 50 µM.
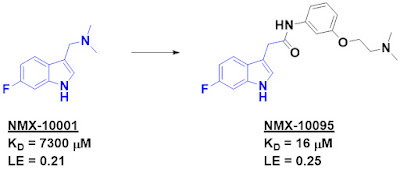
After deconvolution, two hits confirmed, one of which was NMX-10001.
The affinity of the compound was found
to be so low that 1H NMR experiments could not detect binding. Thus,
the researchers kept to fluorine NMR to screen for commercial analogs. They used
19F-detected versions of differential line width (DLW) and CPMG
experiments to rank affinities, and the latter technique was also used to test
for compound aggregation using methodology we highlighted in 2019. Indeed, the researchers
have developed multiple tools for detecting aggregators, such as those we wrote
about in 2022.
Ligand concentrations were measured
by NMR, which sometimes differed from the assumed concentrations. As the
researchers note, these differences, which are normally not measured
experimentally, can lead to errors in ranking the affinities of compounds. The
researchers also examined the 1D spectra of the proteins to assess whether compounds
caused dramatic changes via pathological mechanisms, such as precipitation.
The researchers turned to
protein-detected 2D NMR for orthogonal validation and to determine the binding
sites of their ligands. These experiments revealed that the compounds bind in a
shallow pocket that has previously been targeted by several groups (see here for
example). Optimization of their initial hit ultimately led to NMX-10095, which
binds to the protein with low double digit micromolar affinity. This compound
also blocked SOS-mediated nucleotide exchange and was cytotoxic, albeit at high
concentrations.
I do wish the researchers had
measured the affinity of their molecules towards other RAS isoforms as this
binding pocket is conserved, and inhibiting all RAS activity in cells is
generally toxic. Moreover, the best compound is reminiscent of a series reported
by Steve Fesik back in 2012.
But this specific example is less important
than the clear description of an NMR-heavy assay cascade that weeds out
artifacts in the quest for true binders. The strategy is reminiscent of the “validation cross” we mentioned back in 2016. Perhaps someday computational methods will
advance to the point where “wet” experiments become an afterthought. But in the
meantime, this paper provides a nice set of tools to find and rigorously
validate even weak binders.