The year is spinning to an end, and as we did in 2013 and 2012, Practical Fragments is looking back on
notable events as well as reviews we didn’t cover previously.
2014 was full of conferences, starting with the CHI meeting
in San Diego (here and here), moving to the Zing conference in the Dominican
Republic, on to the Fall ACS meeting in beautiful San Francisco, and ending
with FBLD 2014 in Basel.
In terms of reviews relevant to the fragment community, John
Christopher and colleagues at Heptares published an extensive analysis of “Structure-based and fragment-based GPCR drug discovery” in ChemMedChem
early in 2014. The last few years have seen an efflorescence of new structural
information on G protein-coupled receptors, and this paper provides a thorough
compilation of crystal structures and small molecule ligands. The review also
discusses methods that have been used to discover fragments that bind to GPCRs,
including TINS, SPR, CEfrag, radioligand binding, and fluorescence assays, and
ends with case studies on A2A antagonists and β1AR
ligands.
In contrast to GPCRs, kinases represent a well-established target class for fragment-based drug discovery, as exemplified by the first approved drug, vemurafenib. Structural biology has played a major role in this success; more than 200 of the 518 human kinases have had their X-ray crystal structures determined, and more than 3000 protein kinase structures have been deposited in the protein data bank. Astex has put several kinase inhibitors into the clinic, and in Methods in Enzymology Paul Mortenson and colleagues from the company discuss the state of the art. This is a clear and concise review of fragment-based drug discovery in general and as specifically applied to kinases. It serves as an excellent introduction to the topic.
Any chemist who has worked on kinases will be familiar with
azaindoles, and in Molecules, Sylvain
Routier and colleagues at Université d’Orléans discuss “the azaindole framework in the design of kinase inhibitors.” This provides a thorough compilation of
azaindole inhibitors against ALK, Aurora, Cdc7, CHK1, C-Met, DYRK1A, FAK, IKK2,
JAK2, KIT/FMS, PAK1, p38α, PIM1, B-Raf, ROCK, m-TOR, and TrkA, replete with
synthetic methods. The paper also includes a nice analysis of binding modes. Of
the 58 crystal structures of azaindoles bound to kinases in the protein data
bank, the majority (48) are with 7-azaindole rather than the three other
positional isomers. This isomer (found in vemurafenib) is also over-represented
in the patent literature and among commercial compounds.
Another target that has yielded to FBLD is BACE1, a hot but
still controversial target for Alzheimer’s disease, and in Bioorg. Med. Chem. Lett. Daniel Oehlrich and colleagues at Janssen
review “the evolution of amidine-based brain penetrant BACE1 inhibitors”. This
is very much a medicinal chemist’s review, with over 100 chemical structures,
including a nice summary of the various chemotypes used by different companies.
The authors do an excellent job synthesizing a tremendous amount of data, much
of it reported only in the patent literature, and engage in some intriguing
chemical sleuthing to guess at the identity of clinical candidates whose
structures have not been publicly disclosed, such as MK-8931.
Jia Zhou and collaborators at the University of Texas
Galveston and Fuzhou University discuss “Evolutions in fragment-based drug design: the deconstruction-reconstruction approach” in Drug Discovery Today. After briefly describing fragment-finding
methods and library design, the review focuses on deconstruction of known
ligands to generate “privileged” fragments that are then reassembled into new
molecules. Although this approach can be productive, if one doesn’t exclude
PAINS the result can be garbage-in, garbage-out.
Finally, in Methods in
Enzymology, Katherine Warner (National Heart, Lung and Blood Institute) and
Adrian Ferré-D’Amaré (University of Cambridge) review the crystallographic
analysis of fragments binding to the TPP riboswitch. This is a concise how-to
guide, and the methodology could be applicable to other RNA targets.
And with that, Practical
Fragments says farewell to 2014. Thanks for reading, and may the New Year
bring wonderful new discoveries!






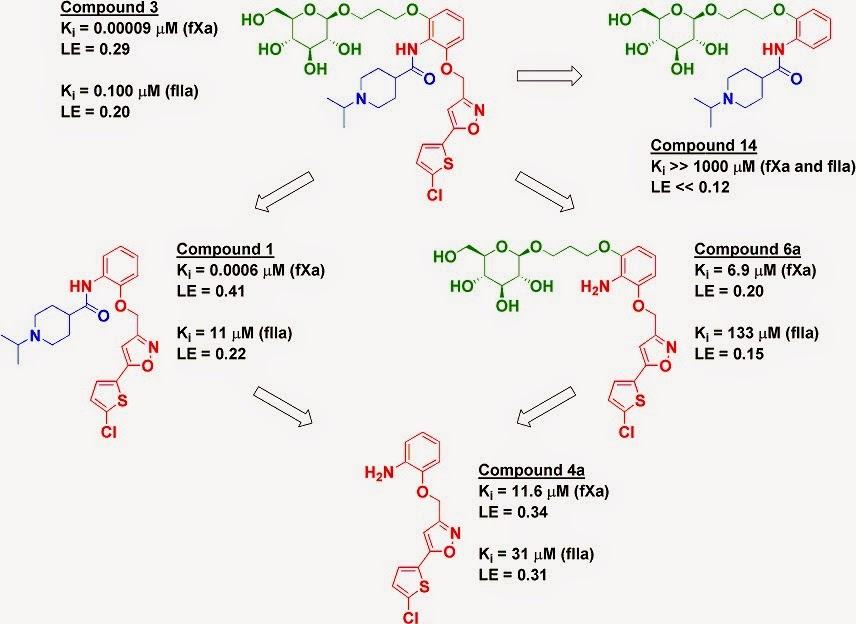



 So, in the end, I was pleasantly surprised. This ended up being nice work with a good breadth of work. I don't know if this makes it into the "lead-like" space or will remain in the "tool" space, but I like to see this kind of work, especially from academic groups.
So, in the end, I was pleasantly surprised. This ended up being nice work with a good breadth of work. I don't know if this makes it into the "lead-like" space or will remain in the "tool" space, but I like to see this kind of work, especially from academic groups. 

