The kinase CK2 is an intriguing anti-cancer target, but most
of the reported inhibitors bind in the conserved hinge region and so also hit
other kinases, complicating interpretation of the biology. A team based at the
University of Cambridge has taken a fragment-linking approach to discover more
selective inhibitors. The first report was published last year by Marko
Hyvönen, David Spring, and colleagues in Chem.
Sci., and they have now published a more complete account in Bioorg. Med. Chem.
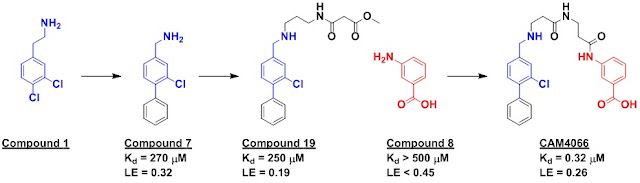
A crystallographic screen identified compound 1, which bound
to six different sites! One of these sites was particularly interesting as it appeared
to be a previously undiscovered “αD” pocket near the ATP-binding site. A couple
cycles of SAR by catalog, informed by computational screening, led to compound
7, which binds in the desired pocket but not at other sites.
Although compound 7 has measureable affinity for CK2α as
judged by ITC, it does not inhibit the enzyme, which is not surprising because
it does not bind in the ATP-binding site. Thus, the researchers screened 352
fragments from Zenobia in cocktails of 4, each at 5 mM, and found 23 that bound
in the ATP site. Reasoning that the hinge region is the most conserved portion
of the ATP-binding site, the researchers avoided fragments that bound there.
This led them to focus on compound 8, which has a synthetic handle pointing
towards the αD pocket.
Next, modeling was used to generate a series of appendages
from compound 7 to try to reach compound 8. Compound 19 looked like it could
bridge the gap, a hypothesis which was confirmed when linking led to a low
micromolar binder. Tweaking the linker led to CAM4066, which showed nanomolar
binding as well as inhibition of CK2. Crystallography revealed that the linked
molecule bound as expected.
CAM4066 was tested against 52 other kinases at 2 µM and showed at
most only 20% inhibition, suggesting that it is indeed quite selective for CK2.
Unfortunately, perhaps because of its carboxylic acid, it did not show any cell
activity. This was addressed by making a methyl ester prodrug – a strategy that
was also taken for another fragment linking campaign on a very different
target.
As the researchers point out, CAM4066 follows the Evotec
model of a largely lipophilic fragment linked to a more polar fragment. There
is still much more to do: no pharmacokinetic data are provided, and the potency
still falls short of what is needed for a chemical probe. Still, this is a nice
illustration of the power of fragment linking, guided by both modeling and
crystallography, to generate molecules with interesting properties.



No comments:
Post a Comment