Of all the ways to find fragments, one of the more unusual
is substrate activity screening, or SAS, which we first discussed here. The
idea is to make and screen libraries of potential enzyme substrates and transform the best ones into inhibitors. In a new paper in J.
Am. Chem. Soc., Jon Ellman and coworkers at Yale University describe how
they used SAS to discover irreversible inhibitors of protein arginine deiminase
3 (PAD3), a potential target for neurodegenerative diseases.
The four human PADs (conveniently named PAD1-4) transform
arginine residues in proteins to citrulline residues, with subtypes distributed
differently across different tissues. The researchers started by making a
library of more than 200 fragment-sized guanidines (the unique side-chain
moiety in arginine) as potential substrates. These were then screened in a
colorimetric assay. Several compounds were found to be processed by the enzyme,
though all were very weak substrates (Km > 10 mM).
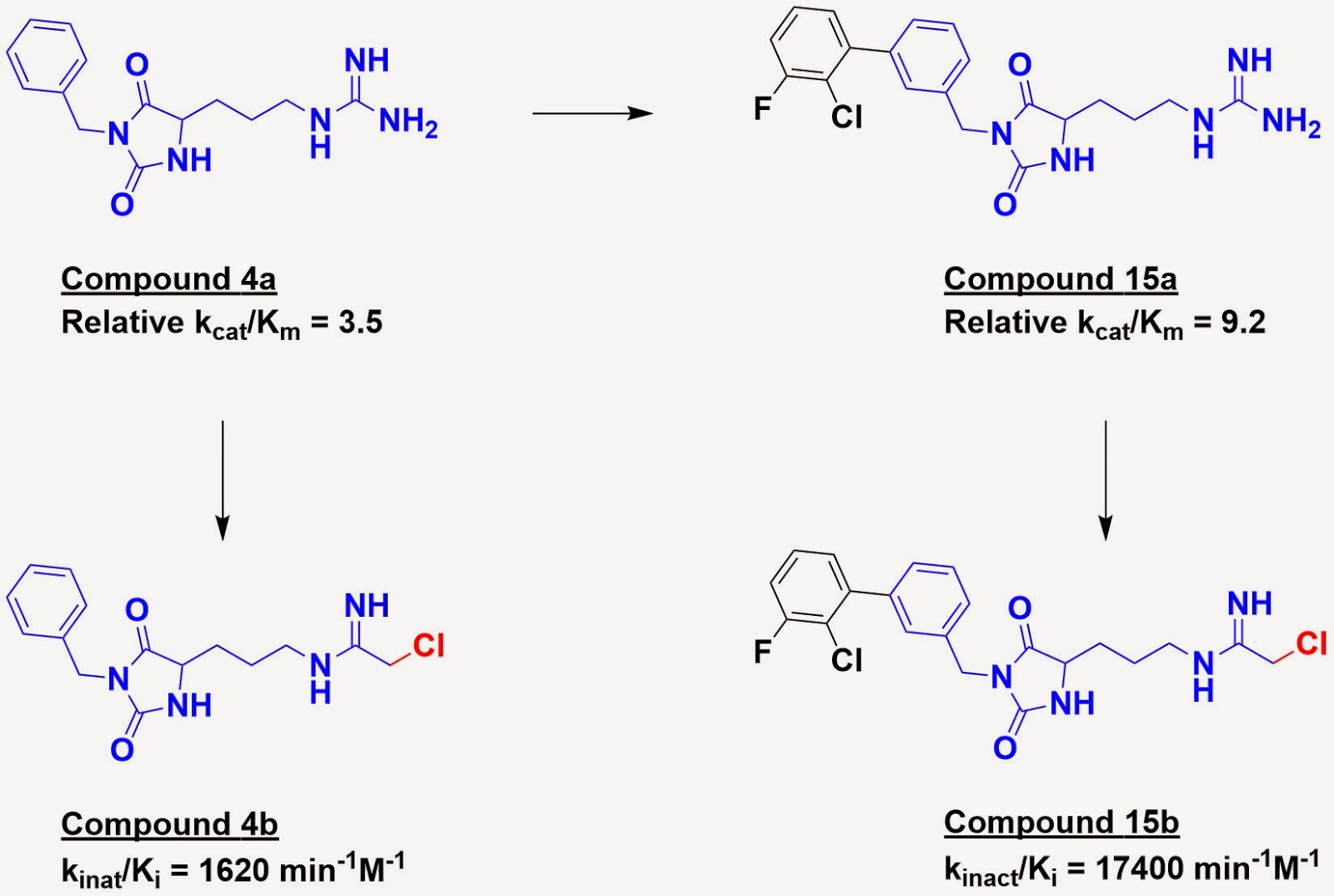
Next, the best substrates from three different chemical
series were optimized for activity. For example, substrate 4a was grown to
substrate 15a.
Finally, the substrates were converted to irreversible inhibitors
by replacing the guanidine with a known chloroacetamide warhead. This coopts
the natural mechanism of the enzyme, which relies on covalent bond
formation between an active-site cysteine residue and the substrate. Within a given series, the better the substrate, the better the
resulting inhibitor (for example, inhibitor 15b is more potent than inhibitor
4b). However, these correlations did not hold across series.
The best compounds were also tested for selectivity, and
some of them were at least 10-fold selective for PAD3 over the other three
PADs.
Last year we highlighted a paper that described several
difficulties encountered (and overcome) using SAS to find inhibitors of the protease urokinase. (The
comments to that post are well worth reading as they include contributions from
the corresponding author of the paper as well as a former Ellman postdoc who is
using SAS.) However, according to the current paper, SAS was relatively straightforward for PAD3 –
another confirmation that different targets require different approaches.



No comments:
Post a Comment