The mitogen-activated protein
kinase (MAPK) signaling pathway is a rich source of targets, particularly for
inflammation. Within this cascade the p38 kinases have been heavily studied,
but many of the inhibitors that entered the clinic derailed for various reasons,
including efficacy. Thus, some groups have sought to block the pathway upstream
of p38. A paper just published online in Bioorg.
Med. Chem. Lett. by Steve Swann and colleagues at Takeda describes some of
their efforts to accomplish this.
The researchers focused on MKK3
and to a lesser degree the related MKK6, both of which phosphorylate and
activate p38. They began by screening their 11,012 fragments in a biochemical
assay at 100 µM each. Hits were prioritized by estimating the IC50
values and thus approximate ligand efficiency (LE) and lipophilic ligand
efficiency values (LLE) for each compound that inhibited >30%. Of these, 93
gave LE ≥ 0.35 kcal/mol per heavy atom and LLE ≥ 4. (Incidentally, this seems
like a perfectly reasonable use of metrics to triage a large number of
compounds, and the speed and simplicity is a good counterargument to more
complicated proposals.) Some hits were tested using full dose-response curves
to determine actual IC50 values and surface plasmon resonance assays
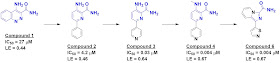
to determine Kd values; compound 1 was particularly compelling.
Readers may recall that Takeda found
this very same fragment as an inhibitor of BTK (a kinase in an unrelated
pathway), and they used the compound/BTK crystal structure along with the
published crystal structure of MKK6 to develop a binding model. In their
pursuit of MKK3/6 inhibitors, the Takeda team performed biochemical screens of
available related compounds. This led to compound 2, which modeling predicted
would bind in a similar fashion. The binding model also suggested the
possibility of picking up a hydrogen bond to a lysine residue, leading to the
more potent compound 3. Further optimization led to compounds 4 and 6, both
with low nanomolar potency against MKK3 and low micromolar or high nanomolar
cell-based activity. Profiling these against a dozen other kinases within the
p38 signaling pathway revealed good selectivity against all except MKK6.
This is a nice, concise paper
that illustrates how modeling, even without direct structural information, can be used to advance a fragment to low
nanomolar inhibitors, albeit in a well-studied class of targets. It is also another illustration that the same fragment can be used to develop completely different series. And finally, these molecules look promising as chemical probes and possibly drug leads; it will be fun to
watch as more data are disclosed.
No comments:
Post a Comment